Achievement
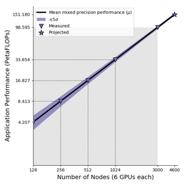
An artificial intelligence system called MENNDL, which used 18,000 NVIDIA Volta GPUs on Oak Ridge National Laboratory's Summit machine, automatically designed an optimal deep learning network in order to extract structural information from raw atomic-resolution microscopy data. MENNDL has been scaled to the 3,000 available nodes of Summit achieving a measured 98.6 PFlops, with an estimated sustained performance of 151 PFlops when the entire machine is available. This work was nominated for the Gordon Bell Award and was selected as one of six finalists.
Significance and Impact
In a few hours, MENNDL creates and evaluates millions of networks using a scalable, parallel, asynchronous genetic algorithm augmented with a support vector machine to automatically find a superior deep learning network topology and hyper-parameter set than a human expert can find in months. For the application of electron microscopy, the system furthers the goal of improving our understanding of the electron-beam-matter interactions and real-time image-based feedback, which enables a huge step beyond human capacity towards nanofabricating materials automatically.
Research Details
- Deep learning code running at 151 PFlops projected on 4600 nodes of Summit.
- Applied to the problem of finding atomic scale defects in materials from scanning transmission electron microscope images.
Overview
Our deep learning framework is called Multinode Evolutionary Neural Networks for Deep Learning (MENNDL). MENNDL relies on two optimization methods, genetic algorithms and support vector machines to intelligently optimize deep learning network topologies and hyperparameters. MENNDL effectively parallelizes network evaluation and fully utilizes the computational power of Summit. The resulting software framework facilitates the discovery of an optimal deep learning network for a particular scientific dataset in a quick, efficient, and automated manner using a GPU-based HPC system.
Using MENNDL, we develop a deep learning network for rapid analysis of dynamic scanning transmission electron microscrope (STEM) data from a 2-dimensional material under electron beam irradiation. This custom network allows us to create a library of defects, map chemical transformation pathways at the atomic level, including detailed transition probabilities, and explore subtle distortions in local atomic environment around the defects of interest. Employing the custom network, we are able to get an unprecedented insight into the nature and mechanisms of solid-state reactions and electron-beam-matter interactions on the atomic level, which is of crucial importance to controllable nanofabrication as well as to fundamental atomic-scale chemistry. Furthermore, the developed network solves the problem of instructing a computer how to choose automatically the "best region" in a sample to make a measurement or perform atomic manipulations without human supervision. This is a critical step towards a fully-automated ("self-driving") microscope.
We utilized the human expert ground truth to train and validate convolutional neural networks using MENNDL. We trained MENNDL for 65 generations and achieved a validation accuracy of 99.51%. We found that MENNDL is able to evolve a network topology and hyperparameter set over time that decreases the validation error, customizing the network to the dataset. The sustained performance for mixed single-half precision is approximately 98.6 PFlops for 3,000 nodes. When Summit's full 4,600 nodes are available, we project that we will achieve approximately 151 PFlops sustained performance for mixed single-half precision. These results were obtained using preproduction system software, not-yet generally available software, and performance results on Summit are expected to improve over coming months.
This material is based upon work supported by the U.S. Department of Energy, Office of Science, Office of Advanced Scientific Computing Research, Robinson Pino, program manager, under contract number DE-AC05-00OR22725.
This research used resources of the Oak Ridge Leadership Computing Facility, which is a DOE Office of Science User Facility supported under Contract DE-AC05-00OR22725.
Last Updated: May 28, 2020 - 4:04 pm